GROUP WITTMANN / Project WI 5350/1-2
Z-Projekt – Bacteriophage Repository, Services and Databank Development
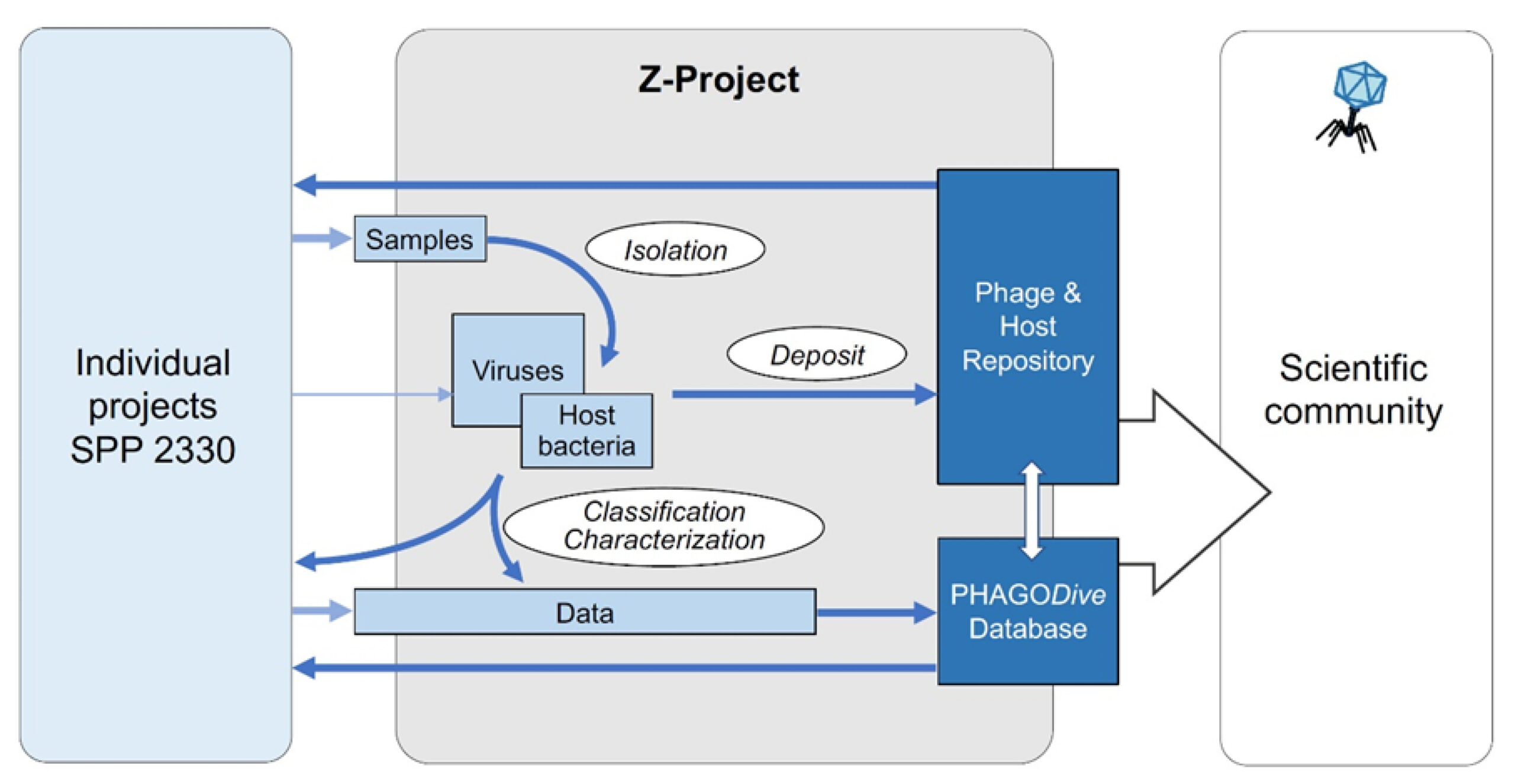
The Z project will provide essential service functions for all participating groups of the SPP and access to state-of-the-art methodologies for studying all relevant aspects of the biology of prokaryotic viruses and will also be in charge of the long-term conservation, quality control and distribution of the isolated bioresources and their data.
Besides isolation of new viruses, the main services within the SPP comprise different aspects of phage biology, including host spectra analyses, growth/lysis kinetics and parameters (burst size, efficiency of plating, etc.), the analysis of phage-host interactions like adsorption at receptors, virus structure via transmission electron microscopy, as well as genomics of both host organisms and bacteriophages.
Besides methodological support of the SPP research projects, one major goal of the Z project conducted at the DSMZ is the development of a new virus-related database, PhageDive. Built on the BacDive model, it gathers and links all the experimental data and metadata (e.g., geographical origin, isolation source) of viruses present in different culture collections. PhageDive aims to facilitate access to all virus information for the scientific community. The database provides information on the taxonomy, host strains, phage morphology, life cycle, origin and genomic data. PhageDive offers advanced search function allowing queries such as „show me all phages infecting this strain“ or „show me all viruses with this enzyme“. The first prototype with the main functions, but limited data has now been launched. In the coming months, data from other culture collections will be integrated and additional features will be developed based on user feedback.
Learn more about PhageDive. Please try it and give feedback:
https://phagedive.dsmz.de
Principal Investigator

Dr. Johannes Wittmann
Leibniz-Institut DSMZ-
Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH
E-Mail: johannes.wittmann@dsmz.de
Homepage: https://www.dsmz.de/research/bioresources-for-bioeconomy-and-health-research/phage-genomics-and-application
Postdoc
Dr. Clara Rolland (Clara.Rolland@dsmz.de)
Publications
- Reimer LC, Vetcininova A, Sardà Carbasse J, Söhngen C, Gleim D, Ebeling C, Overmann J (2019) BacDive in 2019: Bacterial phenotypic data for high-throughput biodiversity analysis. Nucl Acids Res 47(D1):D631-D636.
- Korf IHE, Meier-Kolthoff JP, Adriaenssens EM, Kropinski AM, Nimtz M, Rohde M, van Raaij MJ, Wittmann J (2019) Still Something to Discover: Novel Insights into Escherichia coli Phage Diversity and Taxonomy. Viruses 11(5):454.
