GROUP HÖFER / Project HO 6873/1-1
Understanding bacterial epitranscriptomics on the basis of Escherichia coli – T4 phage interactions
The principal knowledge gap in T4 phage biology is linked to the importance of identifying the molecular mechanisms by which viruses hijack their host’s genetic machinery at the transcriptional and translational level.
In this project, we characterise the bacterial epitranscriptome on the basis of T4 phage – E. coli interaction. By combining next-generation sequencing strategies and proteomic approaches, we will study the molecular influence of post-translational protein modifications such as ADP-ribosylation and RNAylation on the central cellular processes of transcription and translation during phage-host interactions. This study will broaden our understanding of how the ADP-ribosyltransferases of the T4 phage modulate the bacterial key mechanisms and inhibit bacterial defence mechanisms to trigger an effective infection.
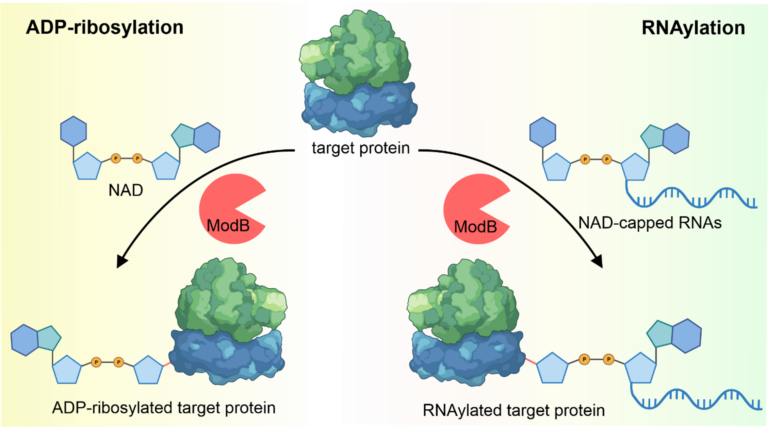
Figure 1: ADP-ribosylation and RNAylation of target proteins by T4 phage ADP-ribosyltransferase ModB. ModB accepts NAD as well as NAD-capped-RNA as substrate, thereby covalently linking ADP-ribose (ADP-ribosylation) or ADP-ribose modified-RNA (RNAylation) to a target protein.
Principal Investigator(s)

Prof. Dr. Katharina Höfer
Max-Planck-Institute for Terrestrial Microbiology
Karl-von-Frisch-Str. 16
35043 Marburg, Germany
E-Mail:katharina.hoefer@synmikro.mpi-marburg.mpg.de
Homepage: https://www.mpi-marburg.mpg.de/856032/Katharina-Hoefer
PhD student(s)
Lydia Garrido Garcia-Dorado (Lydia.Dorado@mpi-marburg.mpg.de)
Publications
- Pozhydaieva, N., Billau, F. A., Wolfram-Schauerte, M., Ramírez Rojas, A. A., Paczia, N., Schindler, D., & Höfer, K. (2024). Temporal epigenome modulation enables efficient bacteriophage engineering and functional analysis of phage DNA modifications. PLoS genetics, 20(9), e1011384.
- Cortés-Martín A, Buttimer C, Pozhydaieva N, Hille F, Shareefdeen H, Bolocan AS, Draper LA, Shkoporov AN, Franz CMAP, Höfer K, Ross RP, Hill C. Isolation and characterization of Septuagintavirus; a novel clade of Escherichia coli phages within the subfamily Vequintavirinae. Microbiol Spectr. 2024 Sep 3;12(9):e0059224. doi: 10.1128/spectrum.00592-24.
- Wolfram-Schauerte, M., Moskalchuk, A., Pozhydaieva, N., Rojas, A. A. R., Schindler, D., Kaiser, S., … & Höfer, K. (2024). T4 phage RNA is NAD-capped and alters the NAD-cap epitranscriptome of Escherichia coli during infection through a phage-encoded decapping enzyme. bioRxiv, 2024-04.
- Pozhydaieva, N., Höfer, K. (2024). Utilizing Golden Gate assembly to streamline CRISPR-Cas/NgTET-based phage mutagenesis (in print in Methods in Molecular Biology, DOI : 10.1007/978-1-0716-4094-4)
- Wolfram-Schauerte, M, Pozhydaieva, N & Höfer, K. Molekulare Schnappschüsse der Phageninfektion (2023) Biospektrum 29, 257–261. https://doi.org/10.1007/s12268-023-1923-x
- Wolfram-Schauerte M, Höfer K. NAD-capped RNAs – a redox cofactor meets RNA (2023) Trends in Biochemical Sciences, 48(2):142-155 DOI: 10.1016/j.tibs.2022.08.004
- Wolfram-Schauerte M, Pozhydaieva N, Viering M, Glatter T, Höfer K. (2023) Integrated Omics Reveal Time-Resolved Insights into T4 Phage Infection of E. coli on Proteome and Transcriptome Levels. Viruses. 12;14(11):2502. doi: 10.3390/v14112502
- Höfer K*, Schauerte M, Grawenhoff J, Wulf A, Welp LM, Billau FA, Urlaub H, Jäschke A* (2021) Viral ADP-ribosyltransferases attach RNA chains to host proteins. bioRxiv, doi: https://doi.org/10.1101/2021.06.04.446905
- Schauerte M, Pozhydaieva N, Höfer K* (2021) Shaping the Bacterial Epitranscriptome—5′-Terminal and Internal RNA Modifications. Advanced Biology, doi: https://doi.org/10.1002/adbi.202100834
