GROUP KAMP / Associated Project
Inference of prokaryotic virus host interactions and emergent dynamics
The knowledge of prokaryote virus biology includes many unknowns, in particular regarding the interactions prokaryotic viruses establish with their microbial environments and the resulting ecological dynamics. Mathematical and computational modelling offers excellent tools for the discovery and description of still new concepts and mechanisms in prokaryotic virus-host interactions.
The temporal evolution of viral and prokaryotic host population sizes can be seen as a kinetic fingerprint. Its characteristics encode underlying interactions ranging from synergy to competition between viruses and their hosts in a variety of ecosystems such as intestinal, aquatic and soil ecosystems. Mathematical and computational modelling are used to decode this information about underlying interactions, to make predictions about system dynamics in different ecological settings and to serve as a tool for hypothesis generation. This allows to move forward from descriptive to predictive ecology of prokaryotic viruses and their hosts.
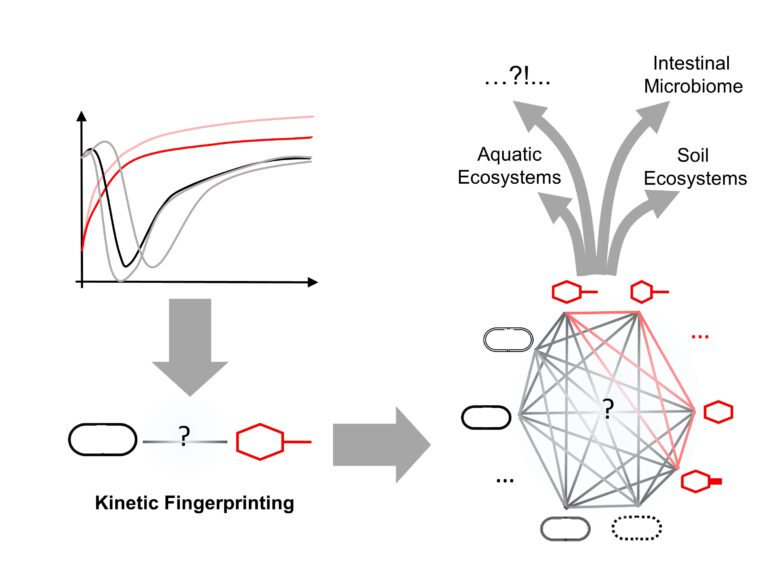
Figure 1: Kinetic fingerprinting is a tool to infer virus host interactions from their population dynamics. Mathematical and computational modelling is used to derive emergent dynamics of larger prokaryotic virus-host communities.
Principal Investigator(s)

PD Dr. Christel Kamp
Paul-Ehrlich-Institut, Langen
Adjunct Lecturer for Bioinformatics
Goethe University Frankfurt
E-Mail: christel.kamp@pei.de
Homepage: https://www.uni-frankfurt.de/91092202/Adjuncts
PhD student(s)
TBA
Publications
Loessner, H., Schlattmeier, I., Anders-Maurer, M., Bekeredjian-Ding, I., Rohde, C., Wittmann, J., Pokalyuk, C., Krut, O.,Kamp, C. (2020) Kinetic Fingerprinting Links Bacteria-Phage Interactions with Emergent Dynamics: Rapid depletion of Klebsiella pneumoniae Indicates Phage Synergy Antibiotics 9(7), 408, https://doi.org/10.3390/antibiotics9070408